Lung cancer, the leading cause of cancer death, is usually diagnosed at a late stage when the survival rate is extremely low. Early-stage lung cancer is mostly asymptomatic, and low-dose spiral CT imaging, the current method for detecting early lung cancer lesions, isn’t feasible as a widespread screening test for the general population due to high cost and the radiation hazard of repeated screenings. A new study, using high-resolution magic angle spinning (HRMAS) magnetic resonance spectroscopy, provides proof-of-concept for the ability of a drop of blood to reveal lung cancer in asymptomatic patients. The study was co-led by researchers at Massachusetts General Hospital (MGH): Leo Cheng and David Christiani.
“Our study demonstrates the potential for developing a sensitive screening tool for the early detection of lung cancer”, says Cheng. “The predictive model we constructed can identify which people may be harbouring lung cancer. Individuals with suspicious findings would then be referred for further evaluation by imaging tests, such as low-dose CT, for a definitive diagnosis.”
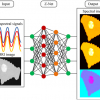
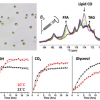
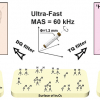
Cheng, Christiani and their co-investigators built a lung-cancer predictive model based on metabolomics profiles in blood. The presence of lung cancer, with its altered physiology and pathology, can cause changes in the blood metabolites produced or consumed by cancer cells in the lungs. The researchers measured metabolomics profiles in blood using HRMAS magnetic resonance spectroscopy. The investigators screened tens of thousands of blood specimens stored in Massachusetts General Hospital’s biobank and others and found 25 patients with non-small cell lung cancer (NSCLC) with stored blood specimens obtained at the time of their diagnosis and at least six months prior to their diagnosis. They matched these patients with 25 healthy controls.
The researchers first trained their statistical model to recognise lung cancer by measuring metabolomic profile values in blood samples obtained from patients at the time of their diagnosis and comparing them to blood samples from the healthy controls. They then validated their model using blood samples from the same patients obtained prior to their lung cancer diagnosis. Here the predictive model yielded values between the healthy controls and the patients at the time of their diagnosis. “This was very encouraging, because screening for early disease should detect changes in blood metabolomic profiles that are intermediate between healthy and disease states”, says Cheng. The investigators then tested their model with a different group of 54 patients with NSCLC using blood samples obtained before their cancer diagnosis, which confirmed that the model’s predictions were accurate.
Values from the predictive model measured from prior-to-diagnosis blood samples could also predict five-year survival for patients, which may be useful in guiding clinical strategies and treatment decisions. A previous study by the investigators showed the potential for magnetic resonance spectroscopy-based metabolomics to differentiate cancer types and stages of diseases. Larger studies are needed to validate the use of blood metabolomics models as NSCLC early screening tools in clinical practice.
Next, the researchers will analyse metabolomic profiles of lung cancer’s clinical characteristics to understand the entire metabolic spectrum of the disease, which may be useful in choosing targeted therapies. They have also measured metabolomics profiles of more than 400 patients with prostate cancer to create a model that will distinguish between indolent cancer, which needs to be monitored, and more aggressive cancer that requires immediate treatment. The investigators also plan to use the same technology to screen for Alzheimer’s disease using blood samples and cerebrospinal fluid.