Colette Germon,a Antony N. Daviesb,c and Paul Jonesa
and Paul Jonesa
aAnalytical and Forensic Science, School of Applied Sciences, Faculty of Computing, Engineering and Science, University of South Wales, UK
bStrategic Research Group – Measurement and Analytical Science, Akzo Nobel Chemicals b.V., Deventer, the Netherlands
cSERC, Sustainable Environment Research Centre, Faculty of Computing, Engineering and Science, University of South Wales, UK
With numerous food-fraud cases being revealed by the media and catching significant public attention, a training project which embodies the establishment of “facts” amongst all the hype is a very good idea. Intergovernmental regulations also control fish catch quotas, which has led to some fraudulent mislabelling of cheaper fish, which then also brings the accuracy of the fish catch data and subsequent fisheries management into doubt. Rapid, simple and robust analytical spectroscopic techniques have the potential to unmask these food fraudsters. Recently, I have enjoyed being involved on the edges of a very nicely crafted and fun experimental project which cleverly brings together learning goals around analytical science, more advanced spectroscopic training, as well as fundamental understandings around food authentication and the applicability of chemometrics techniques.
The problem posed
As part of an MSc project, a challenge was set to see how attenuated total reflection–infrared spectroscopy (ATR–IR) could be used as part of a simple testing methodology for adulteration of food. The experimental design looked not only at mislabelling the source of meat or fish, but was also extended to investigate if the technique could help identify the selling of frozen and then thawed food products as “fresh”. One of the teaching goals is, of course, that you need to sample as widely as possible if you are going to fully cover the scope of the problem. However, this is after all a teaching opportunity rather than a full-scale national screening programme and the lessons learnt would be equally applicable at national or international level.
Representative meat samples chosen
The size of the experimental design can be very nicely scaled to the amount of laboratory instrument time available at the teaching location, as most of the lessons learnt will be valid regardless of the breadth of samples sourced. As several months were available in the current course example, a wide range of meats and fish were added to the design. Processed foods are tough products to clearly characterise and the project started off on pork (shoulder) and beef (rump) samples from a reputable local source of traceable meat. The raw materials were made up into experimental beef burgers of varying mixture concentrations to simulate label fraud from pure beef to pure pork in 10% increments. Two different locations on each sample burger were sampled and three repeat measurements were taken at each location.
These measurements allowed the student to look first for isolated peaks which might be capable of being quantified (not possible) and then to realise the value of chemometric analyses in classifying adulterated vs non-adulterated and creating a regression model for the degree of adulteration. This was a nice combination of practical manual laboratory sample preparation work, analytical spectroscopy using a modern simple accessory and more advanced data analysis applications.
For “unknown” samples, luxury-branded, high-end supermarket outlet burgers were taken and analysed using the same methodology of two locations on the burger and three repeat measurements.
The final issue this piece of work was configured to address was the question of frozen then thawed meat potentially entering the food chain under the mislabelling of fresh. Essentially the same experimental design as above was conducted but using meat which had been frozen at –12°C for three weeks, including the supermarket brand products for comparison, yielding 84 samples for analysis.
Something fishy on the spectrometer
Moving on to fish samples allowed a broad range of samples to be analysed whose spectra were at times extremely similar (see Table 1). Some fish samples had undergone “pre-processing”, such as smoking or had other distinguishing claims such as the location of the source on the fish itself—presumably claims which were intended to imply added quality to the labelling of the product and hence increased price.
Table 1
Fish samples | |
Common name | Latin name |
Mackerel | Scomber scomber |
Sea bass | Dicentrarchus labrax |
Salmon | Salmo salar |
Smoked haddock | Melanogrammus aeglefinus |
Cod | Gadus morhua |
Smoked river cobbler | Pangasius hypophthalmus |
Haddock loin | Melanogrammus aeglefinus |
British hake | Merluccius merluccius |
Plaice | Pleuronectes platessa |
Pollock | Pollachius |
Lemon sole | Microstomus kitt |
Sea bream | Sparidae |
Halibut | Hippoglossus hippoglossus |
Keeping with the theme of processed foods being the easiest product type with which to hide food fraud, the fish samples were all homogenised and prepared into fishcakes before undergoing infrared analysis using the same methodology as in the meat case. Again, three shop-bought “cod” fishcakes were used as “unknowns” against which to test. Finally, fish stock samples were also frozen for a period of weeks to test for the fresh vs frozen/thawed storyline and the measurement sequence repeated.
Equipment and software
The spectra were recorded using a Perkin-Elmer UATR Two FT-IR spectrometer with a one-bounce diamond crystal ATR accessory. Data were acquired by accumulating four scans of 500–4000 cm–1 and the background recorded every six spectra. The cell was thoroughly cleaned between samples. As ATR spectra can be pressure sensitive this was tested for the samples in this study at two different pressures and as no noticeable difference was found the lower pressure was selected to reduce sample compression.
The Unscrambler software package from CAMO was used to carry out the chemometric analyses using partial least squares (PLS) for the quantitative analysis and principal component analysis (PCA) for classification. The only pre-processing considered was wavelength selection at four different ranges, 4000–400 cm–1, 4000–1000 cm–1, 4000–2500 cm–1 and the fingerprint region (1850–1000 cm–1). A “leave-one-out” full cross-validation was used to test the models created.
Some meaty results
The PLS analysis of the beef/pork mixtures went well (see, for example, Figure 1). Regardless of the wavelength range selected or whether the frozen/thawed samples were analysed, the R2 values showed good calibrations were obtained, ranging from 0.97 to 0.99. The best results were obtained for the 4000–1000 cm–1 wavelength range with 0.99.
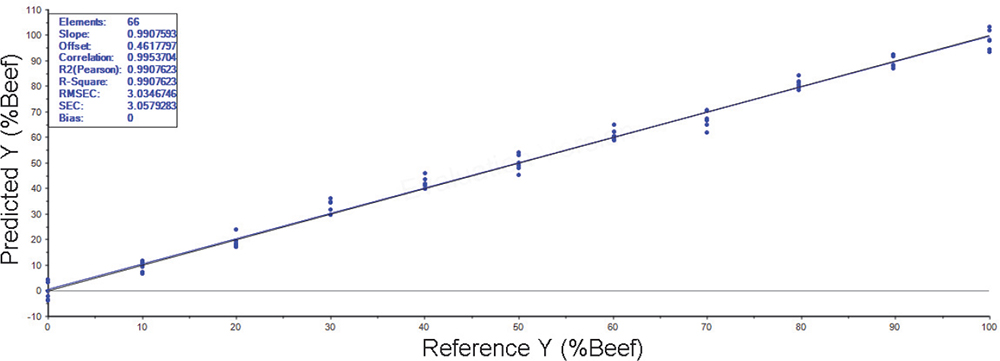
Figure 1. PLS analysis of the fresh meat.
The supermarket burgers when analysed, pleasingly showed good correlation between the label values, however, Figure 2 clearly shows that these burgers contained significantly more fat at the cost of the protein levels than those prepared from the fresh meat.
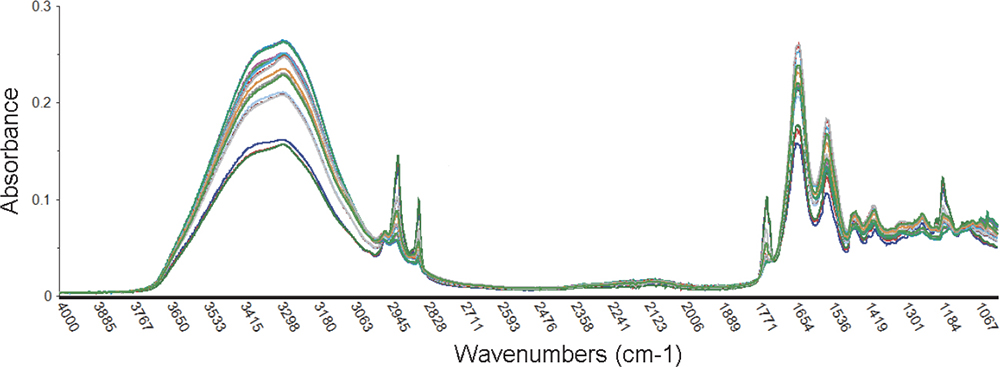
Figure 2. Variation of spectra obtained from the supermarket luxury branded beef burgers at wavelength range 4000–1000 cm–1.
Some fishy results
The study results for the potential fish adulteration also delivered excellent calibration results (e.g. Figure 3 for Cod with Haddock adulteration with R2 value of 0.99). This issue occasionally rises its head in the normal newspapers for public consumption, such as a report on figures supplied by councils to the UK Food Standards Agency indicating that 41 of 303 checks on packaged frozen or chilled fish and in catering businesses did not meet labelling requirements.1 They reported that in 27 cases the fish described as Cod was another species, such as Haddock, Whiting or Pollock. Other types of fish were also incorrectly described among other breaches of labelling rules. In Ireland, one DNA testing study reportedly found that 28% of the fish labelled as Cod from 226 products in 131 supermarkets, fishmongers and takeaways were actually substitutes, such as Pollock and Whiting.
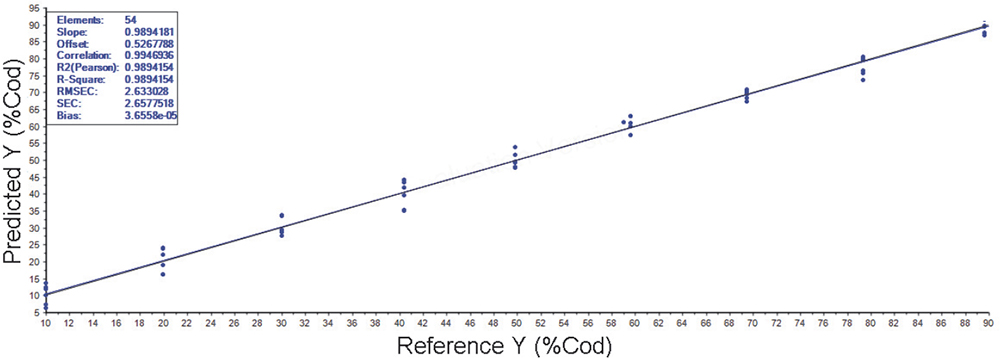
Figure 3. PLS regression for frozen/thawed Cod with Haddock adulteration (10–90%) calculation data.
Figure 4 shows that the infrared spectra of some of the fish species selected are remarkably similar, in this case Cod and Haddock. This makes the classification of species difficult.
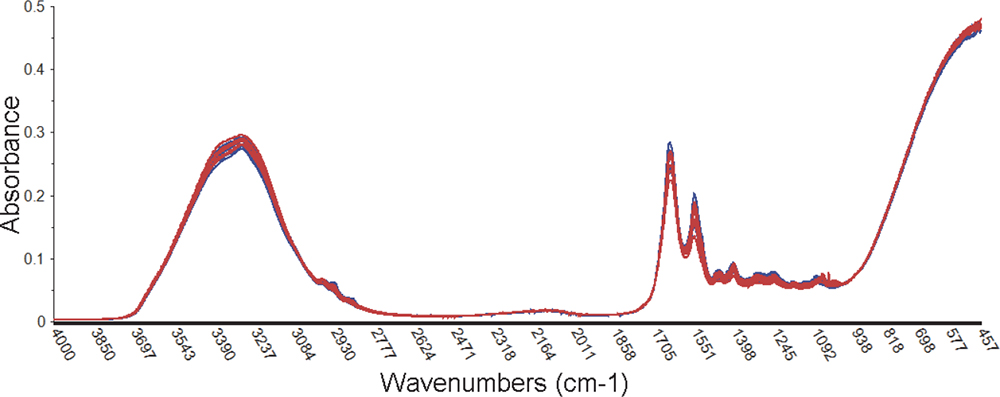
Figure 4. The similarity of the infrared spectra of two fish species is substantially greater than between the two meat sources analysed.
Summary
Before all our food scientists leap to their Facebook and Twitter accounts to decry the breadth of the work done, it is worth reminding ourselves that this is an innovative approach to helping teach and embed understanding in students of some pretty complex and difficult topics. If we look into the literature, one of the interesting challenges in, for example, the area of fish labelling is that the common names for fish vary from location to location. FishBase2,3 is a great source of information in this respect and, when searched, showed that “Cod” was used in the USA alone for six different species, only one of which is the same as used in Europe. Also, there is no species overlap between what you will be served in the USA and Europe and Australia where “Cod” may be two different species, one of which is a Perch.
All in all, I was very impressed with the development of this particular project and its potential future use as a teaching tool for spectroscopic data handling and advanced analysis techniques.
If you have similar examples which we can learn from please get in touch!
References
- https://www.theguardian.com/lifeandstyle/2013/apr/02/fish-chips-cod-food-standards (accessed 23 June 2017).
- http://www.fishbase.org when accessed they claimed to have data covering 33,500 Species, 319,000 Common names, 58,100 Pictures, 53,800 References, 2270 Collaborators and were enjoying 700,000 visits per month!
- R. Froese and D. Pauly (Eds), FishBase, version 02/2017. (2017). www.fishbase.org.