Maria C. Prieto Conaway,a Shousong Cao,b Farukh Durrani,b Youcef Rustum,b Ping Wang,c Khin Marlarc and Latif Kazimc
aThermo Fisher Scientific, San Jose, CA, USA
bRoswell Park Cancer Institute, Department of Cancer Biology, Buffalo, NY, USA
cRoswell Park Cancer Institute, Department of Cell Stress Biology, Buffalo, NY, USA
Introduction
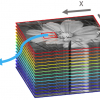
The majority of diseases that lead to premature death in humans originate in tissue and cell dysfunction, due to either congenital or acquired genetic deficiency. Tissue imaging is a promising technique as it allows scientists to follow, among other things, distribution patterns derived from drug treatment (precursor drug and metabolites) in specific tissues or their distribution within, for example, a whole rat. One of the biggest problems in tissue imaging is the amount of background ”noise”, which can make it very difficult to identify the analyte of interest among all the other signals present. Tissue samples are extremely complex, with many different lipids, sugars, proteins and peptides overwhelming any trace analytes in a spectrum. This article discusses matrix-assisted laser desorption/ionisation (MALDI) enabled linear ion trap (LIT) mass spectrometry (MS) as a technique for fast and accurate tissue imaging, compared to the more traditional time-of-flight (ToF) method.