Marek Šebela
Department of Protein Biochemistry and Proteomics, Centre of the Region Haná for Biotechnological and Agricultural Research, Faculty of Science, Palacký University, Olomouc, Czech Republic. E-mail: [email protected]
Introduction
This review article describes the use of solid mixed matrices in matrix-assisted laser desorption/ionisation (MALDI) time-of-flight (TOF) mass spectrometry (MS). These are prepared by combining common matrix compounds and applied particularly to analyse peptides, proteins, oligosaccharides, oligonucleotides, lipids, synthetic polymers or intact cells. The biggest advantage of this approach is that it typically yields homogeneous sample spots on the target plate and provides highly reproducible measurements accompanied by increased sensitivity and resolution.
Since its discovery towards the end of the 1980s, MALDI, which is usually associated with a TOF mass analyser in commercial instruments, has become a common technology in biological MS.1 MALDI-TOF MS has found its application not only in the analysis of peptides and proteins, oligonucleotides, oligosaccharides, technical polymers and small polar compounds, but also viruses or even intact microbial cells. Initially there was an opinion that it would never be possible to analyse large biomolecules using mass spectrometry. It was assumed that, as first it is necessary to vaporise and then ionise the sample to perform MS, large molecules would be decomposed long before their vaporisation. Moreover, mass spectrometers were originally designed for small molecular masses not exceeding 1 kDa.2
Laser desorption/ionisation for biomolecules was independently introduced by Michael Karas and Franz Hillenkamp (Germany) for solids in a crystalline environment3 and Koichi Tanaka (Japan) in the liquid phase.4 The use of lasers to generate ions dates back to the 1960s when a majority of the initial experiments was done with inorganic compounds. The primary role for the laser was to produce a very rapid heating, and the energy transfer was considered to occur via the metallic substrate.5 Karas and Hillenkamp, who actually coined the term MALDI, first recognised that a small organic compound, which strongly absorbed at the laser wavelength they used and exhibited a soft ionisation, could enhance the ionising laser desorption process. During analyses with amino acids, the highly absorbing tryptophan functioned well as a matrix for alanine, a non-absorbing amino acid, when they were present together in the form of a 1 : 1 mixture.3 Later on, Karas and Hillenkamp worked with nicotinic acid as a matrix with excitation at 266 nm. Having been aware of results of Koichi Tanaka and his coworkers,4 they successfully turned their increased effort with a new post-acceleration detector into a finding of the possibility to ionise and detect proteins with molecular masses above 10 kDa.6 In contrast, Tanaka and his coworkers relied on an ultrafine cobalt powder with glycerol for laser desorption and ionisation and to their satisfaction, they could measure proteins up to about 35 kDa.4 Nowadays, this method is rarely used compared with the approach by Karas and Hillenkamp. Nevertheless, Koichi Tanaka was awarded the Nobel Prize for Chemistry in 2002 together with John Bennett Fenn, who in parallel discovered electrospray ionisation (ESI), another soft ionisation technique, which is also well suited to studying biological macromolecules.2
The role of matrix in MALDI process
Competition between the two soft ionisation techniques saw ESI-MS pushed forward in its development as it offered the advantage of being able to be coupled on-line with high-performance liquid chromatography. In the 1990s, the acceptance of MALDI MS as an analytical tool was raised by introducing both delayed extraction of ions (resulting in high resolution spectra) and post-source decay fragmentation (allowing one to obtain structural information on selected ions).5 The process of MALDI is achieved in two steps.7 First, the analyte to be investigated is typically dissolved in a solvent, which contains an appropriate matrix compound. On the target plate, a “solid-solution” deposit of analyte-doped matrix crystals is formed on evaporation of the solvent. As a result, molecules of the analyte are embedded in an excess of matrix molecules.7 In the second step, which takes place under vacuum conditions in the ion source of the spectrometer, the co-crystals are ablated in portions by intense laser pulses (see Figure 1).

The exact mechanism of the formation of ions is not completely understood. Definitely, the matrix is indispensable in this process. The rapid heating by laser irradiation causes the accumulation of energy through excitation of the matrix molecules. A dense plume of material containing both the matrix and matrix–analyte clusters expands into the vacuum of the ion source.8 The most widely accepted mechanism of primary ionisation involves generation of charged species within a cluster ablation process or gas-phase proton transfer in the expanding plume from photo-ionised matrix molecules.7,8 The ions in the gas phase are then accelerated by an electrostatic field and move towards the analyser.
Interestingly, nearly all of the matrix compounds that are routinely used today were discovered in the early days of MALDI.5 In 1989, Beavis and Chait introduced cinnamic acid derivatives, such as sinapinic, ferulic and caffeic acids (SA, FA, CA, respecively),9 which performed excellently for the analysis of proteins using ultraviolet lasers with maximum wavelengths at 355 nm (Nd : YAG) or 337 nm (nitrogen laser).5 After a few years, the same researchers discovered that a-cyano-4-hydroxycinnamic acid (CHCA) was efficient for MALDI-TOF MS analysis of peptides and glycopeptides in the molecular mass range of 500–5000.10 2,5-Dihydroxybenzoic acid (DHB) has become another matrix suitable for peptides and glycopeptides, which was first examined at that time.11 Currently, there are many compounds available which can be applied for this purpose according to the specific needs, but only a few of them are really good. Anyway, matrix choice and optimisation of the sample preparation protocol are the most important steps in MALDI experiments.7 The final selection is always related to the chemical nature of the analyte and the respective laser wavelength. The most effective MALDI matrix must simultaneously meet a number of general requirements:7 strong absorbance of the laser light at a given wavelength, ability to sublime, vacuum stability, promotion of analyte ionisation, solubility in analyte-compatible solvents and lack of reactivity. The choice of matrix is also important for the control of fragmentation, especially in the case of proteins and peptides carrying post-translational modifications. A small number of compounds have been studied with respect to their propensity to induce fragmentation. They are thereby classified as “hot” (assisting the formation of ions with a high internal energy) or “cold”. For example, Karas et al. found that post-source decay (= metastable fragmentation) decreases in the order CHCA or SA > DHB mixed with 2-hydroxy-5-methoxybenzoic acid (so-called “super DHB”) > 3-hydroxypicolinic acid (HPA) for protonated glycoproteins.12 However, it has been shown that the assignment of “hot” and “cold” matrix may reverse as the analyte or laser energy changes.13
Mixed matrices for proteins, peptides, oligosaccharides and oligonucleotides
Various approaches have been explored to enhance matrix selectivity, such as the use of additives, but they do not directly affect the ionisation and rather influence co-crystallisation or exclusion of salt impurities. A big potential resides in combining more compounds into binary or ternary matrix mixtures. The above-mentioned “super DHB” is a mixture of DHB and 2-hydroxy-5-methoxybenzoic acid in a weight ratio of 9 : 1. This binary matrix provides higher yields and signal-to-noise ratios for analyte ions in the high mass range: typically it is applicable to intact proteins. The reason is elucidated by the formation of a disorder in the DHB crystal lattice resulting in “softer” desorption.14 Another combination based on DHB involves CHCA as the second matrix component (in a weight ratio of 1 : 1).15 This mixed matrix proved to be well suited for peptides and intact proteins (an improved signal intensity and resolution was found especially for glycoproteins). The crystallisation of samples with CHCA/DHB results in a homogeneous pattern of CHCA-derived crystals in the central part of the sample spot with small DHB crystals at the perimeter. When applied, CHCA/DHB provides both significantly improved spot-to-spot reproducibility of mass spectra and reduced noise compared with the use of either component individually, which simplifies fully automated acquisition. An interesting binary matrix for measuring phosphopeptides was discovered by combining HPA with CHCA in an optimised weight ratio of 1 : 1 (see Figure 2). As a result, phosphopeptide signals can be acquired reproducibly in either the positive- or negative-ion mode at a lower laser power than with DHB (a common matrix for phosphopeptides) or CHCA alone. In addition, the analysis time is shortened by avoiding searching for the so-called “sweet” spots as is necessary when working with DHB. Concurrently, neutral losses of phosphate group (–80 Da) and phosphoric acid (–98 Da) are reduced. The mixed matrix is also applicable to intact phosphoproteins (as has been demonstrated with molecules up to 30 kDa).16

Oligosaccharides represent a challenge for MALDI-TOF MS as there are typically many problems with low sensitivity and strong background noise. Neutral oligosaccharides are detected in the positive-ion mode as sodium- or potassium-charged pseudomolecular ions for which DHB usually represents the matrix of choice. Oligosaccharides from human milk have been studied with numerous matrices, their binary mixtures and single-matrix mixtures with various additives.17 DHB with 2,4-dinitrobenzoic acid and a diluted NaCl solution was found optimal for standard analyses with neutral compounds. 5-Chloro-2-mercaptobenzothiazole proved to be an excellent first layer for DHB and 6-aza-2-thiothymine (ATT; by the way this is perfect for investigations on acidic oligosaccharides in the negative-ion mode) or could be used alone.17 Combining DHB with the basic aminopyrazine in a weight ratio of 3 : 1 allowed an excellent data acquisition not only for neutral monosaccharides and oligosaccharides (including the content of a real plant extract) but also maltodextrins up to 40 glucose units.18 In contrast, the presence of salts is undesirable when performing MALDI-TOF MS with oligonucleotides because of the formation of adducts between the phosphate backbone of DNA and counter ions (Na+, K+). Nowadays, oligonucleotides are largely utilised as gene probes in molecular biology. When the quality evaluation of synthetic preparations by MALDI-TOF MS is necessary, HPA, ATT or 2¢,4¢,6¢-trihydroxyacetophenone are suitable matrices.7 Diammonium hydrogen citrate is used as a common additive. A major obstacle is the relatively poor mass resolution, which increases in parallel with the increasing molecular mass accompanied by the decrease in signal intensity and sensitivity. To cope with these troubles, the use of a mixture of HPA with pyrazinecarboxylic acid (4 : 1, v/v, for saturated solutions) has been described, which provides better spectral parameters as well as higher reproducibility and tolerance to impurities.19
Mixed matrices for lipids, low-molecular-weight compounds, polymers and intact microbial cells
Dithranol (1,8-dihydroxy-9,10-dihydroanthracen-9-one) is recommended for MALDI-TOF MS experiments with lipids.7 A ternary matrix containing dithranol, DHB and CHCA (in hexane/isopropanol/dimethylsulfoxide; mixed in an equimolar ratio with the addition of sodium iodide) has recently been employed to analyse lipid impurities in biodiesel, which is composed mainly of fatty acid methyl esters.20 The biggest advantage of the ternary matrix mixture resided in the formation of smaller and more homogeneous co-crystals with samples (compared with the use of individual matrices), which had a positive impact on reproducibility and sensitivity. Moreover, semi-quantitative information could be obtained using a calibration with standards. Similarly, a binary matrix consisting of CHCA and DHB in a weight ratio of 1 : 1 in 70% methanol/0.1% TFA (trifluoroacetic acid) containing 1% piperidine was found efficient for MALDI imaging MS of phospholipids.21 The binary mixture was deposited onto a sliced rat brain tissue and, importantly, it crystallised homogeneously, which resulted in increased reproducibility and signal intensities as well as a higher number of signals. The matrix was superior to conventional matrices in its performance and functioned well in both positive- and negative-ion modes: more than 100 glycerophospholipids and sphingophospholipids could be identified by MS and tandem MS.21 As a certain disadvantage, a strong background at masses below 500 Da appeared which fortunately did not disturb the use of CHCA/DHB for phospholipids. The problem of background signals at low masses was addressed by Guo and He22 in 2007: a binary matrix combining CHCA and basic 9-aminoacridine (the compounds differ from each other in their affinity to protons) significantly reduced the number of background matrix peaks in both positive- and negative-mode detection of small molecules. In addition, better signal-to-background ratios were observed for negatively charged inositol phosphates in complex plant extracts. Various binary matrices (e.g. DHB + CHCA, DHB + SA, CHCA + SA and others based on 2,6-dihydroxybenzoic acid as an isomer of DHB) or ternary mixtures such as DHB + CHCA + SA or 2,6-dihydroxybenzoic acid + CHCA + SA allowed one to achieve an improved sensitivity in the analysis of polyethylene glycols with molecular masses ranging from 1 kDa to 10 kDa (the reason probably resides in more homogeneous matrix-sample co-crystal patterns).23
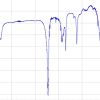
FA is a commonly used matrix for intact cell/spore MALDI-TOF MS of fungi. In respect of the number of ion species and signal intensities, FA was shown to be superior, for example, to CHCA, SA, 2-hydroxy-5-methoxybenzoic acid or 2¢,4¢,6¢-trihydroxyacetophenone when Fusarium species were analysed despite involving additives or applying binary matrix combinations such as DHB/2-hydroxy-5-methoxybenzoic acid and CHCA/DHB.24 One major drawback of FA is the inhomogeneous look of its co-crystals with the analyte resembling snow-covered branches. In our laboratory, we have a long and extensive experience in the use of binary matrices for intact fungi. To find the optimal matrix, CA, CHCA, DHB plus FA were evaluated individually and then also in the form of various binary mixtures such as CHCA/FA, CHCA/SA and FA/SA (different mixing ratios and solvents). The mixture of FA and SA in a weight ratio of 1 : 3 (specifically 5 : 15 mg mL–1) in acetonitrile/2.5% (v/v) TFA, 7 : 3, v/v, was finally proven to be optimum for working with fungi and oomycetes.25 It allows a reproducible acquisition of signals (because of the homogeneous crystallisation pattern) with relatively high signal-to-noise ratios, which are distributed evenly over a large mass region (see Figure 3). Also a binary mixture CA/SA or ternary mixture with CA, FA and SA represent good choices for working with fungal cells (unpublished results).

Conclusions
The preparation of samples for MALDI-TOF MS is an exciting experimental chemistry. The choice of the best matrix, solvents and sample preparation technique are crucial steps on the way to achieving reliable results. To overcome certain imperfections in the use of common matrix compounds, there is a possibility of applying additives or making binary or ternary matrix systems. Combining more compounds into a final mixture allows optimisation of the different properties required of the matrix by varying the identity and/or concentration of components. As a consequence, the sample preparation step provides more homogeneous crystallisation on the target plate, which ensures higher reproducibility of measurements and is typically accompanied by both higher sensitivity and better resolution of signals.
Acknowledgements
This work was supported by grant no. L01204 from the Ministry of Education, Youth and Sports, Czech Republic (the National Program of Sustainability I).
References
- A. Croxatto, G. Prod’hom and G. Greub, “Applications of MALDI-TOF mass spectrometry in clinical diagnostic microbiology”, FEMS Microbiol. Rev. 36, 380 (2012). doi: http://dx.doi.org/10.1111/j.1574-6976.2011.00298.x
- J. Franzen, “ Mass spectrometry in Bremen, a tribute to Dr Ludolf Jenckel”, in A History of European Mass Spectrometry, Ed by K.R. Jennings. IM Publications, Chichester, p. 152 (2012).
- M. Karas, D. Bachmann and F. Hillenkamp, “Influence of the wavelength in high-irradiance ultraviolet laser desorption mass spectrometry of organic molecules“, Anal. Chem. 57, 2935 (1985). doi: http://dx.doi.org/10.1021/ac00291a042
- K. Tanaka, H. Waki, Y. Ido, S. Akita, Y. Yoshida and T. Yoshida, “Protein and polymer analyses up to m/z 100,000 by laser ionization time-of-flight mass spectrometry“, Rapid Commun. Mass Spectrom. 2, 151 (1988). doi: http://dx.doi.org/10.1002/rcm.1290020802
- M. Karas, “From field desorption to MALDI and to the resurgence of time-of-flight mass spectrometry” in A History of European Mass Spectrometry, Ed by K.R. Jennings. IM Publications, Chichester, pp. 70–95 (2012).
- M. Karas and F. Hillenkamp, “Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons“, Anal. Chem. 60, 2299 (1988). doi: http://dx.doi.org/10.1021/ac00171a028
- E. de Hoffmann and V. Stroobant, “Ion sources”, Mass Spectrometry Principles and Applications. John Wiley & Sons, Chichester, pp. 33–39 (2007).
- M. Karas and R. Krüger, “Ion formation in MALDI: The cluster ionization mechanism“, Chem. Rev. 103, 427 (2003). doi: http://dx.doi.org/10.1021/cr010376a
- R.C. Beavis, B.T. Chait and H.M. Fales, “Cinnamic acid derivatives as matrices for ultraviolet laser desorption mass spectrometry of proteins“, Rapid Commun. Mass Spectrom. 3, 432 (1989). doi: http://dx.doi.org/10.1002/rcm.1290031207
- R.C. Beavis, T. Chaudhary and B.T. Chait, “a-Cyano-4-hydroxycinnamic acid as a matrix for matrix-assisted laser desorption mass spectrometry“, Org. Mass Spectrom. 27, 156 (1992). doi: http://dx.doi.org/10.1002/oms.1210270217
- K. Strupat, M. Karas and F. Hillenkamp, “2,5-Dihydroxybenzoic acid: a new matrix for laser desorption—ionization mass spectrometry“, Int. J. Mass Spectrom. Ion Proc. 111, 89 (1991). doi: http://dx.doi.org/10.1016/0168-1176(91)85050-V
- M. Karas, U. Bahr, K. Strupat, F. Hillenkamp, A. Tsarbopoulos and B.N. Pramanik, “Matrix dependence of metastable fragmentation of glycoproteins in MALDI TOF mass spectrometry“, Anal. Chem. 67, 675 (1995). doi: http://dx.doi.org/10.1021/ac00099a029
- G. Luo, I. Marginean and A. Vertes, “Internal energy of ions generated by matrix-assisted laser desorption/ionization“, Anal. Chem. 74, 6185 (2002). doi: http://dx.doi.org/10.1021/ac020339z
- M. Karas, H. Ehring, E. Nordhoff, B. Stahl, K. Strupat, F. Hillenkamp, M. Grehl and B. Krebs, “Matrix-assisted laser desorption/ionization mass spectrometry with additives to 2,5-dihydroxybenzoic acid“, Org. Mass Spectrom. 28, 1476 (1993). doi: http://dx.doi.org/10.1002/oms.1210281219
- S. Laugesen and P. Roepstorff, “Combination of two matrices results in improved performance of MALDI MS for peptide mass mapping and protein analysis”, J. Am. Soc. Mass Spectrom. 14, 992 (2003). doi: http://dx.doi.org/10.1016/S1044-0305(03)00262-9
- L.H. Zhou, G.Y. Kang and K.P. Kim, “A binary matrix for improved detection of phosphopeptides in matrix-assisted laser desorption/ionization mass spectrometry”, Rapid Commun. Mass Spectrom. 23, 2264 (2009). doi: http://dx.doi.org/10.1002/rcm.4139
- A. Pfenninger, M. Karas, B. Finke, B. Stahl and G. Sawatzki, “Matrix optimization for matrix-assisted laser desorption/ionization mass spectrometry of oligosaccharides from human milk”, J. Mass Spectrom. 34, 98 (1999). doi: http://dx.doi.org/10.1002/(SICI)1096-9888(199902)34:23.0.CO;2-N
- M.A. Hashir, G. Stecher and G.K. Bonn, “Identification of low molecular weight carbohydrates employing new binary mixtures for matrix-assisted laser desorption/ionisation mass spectrometry”, Rapid Commun. Mass Spectrom. 22, 2185 (2008). doi: http://dx.doi.org/10.1002/rcm.3602
- L. Zhou, H. Deng, Q. Deng and S. Zhao, “A mixed matrix of 3-hydroxypicolinic acid and pyrazinecarboxylic acid for matrix-assisted laser desorption/ionization time-of-flight mass spectrometry of oligodeoxynucleotides“, Rapid Commun. Mass Spectrom. 18, 787 (2004). doi: http://dx.doi.org/10.1002/rcm.1404
- C.R. McAlpin, K.J. Voorhees, T.L. Alleman and R.L. McCormick, “Ternary matrix for the matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI–TOF MS) analysis of non-fuel lipid components in biodiesel”, Energy Fuels 25, 5407 (2011). doi: http://dx.doi.org/10.1021/ef201257g
- S.R. Shanta, L.H. Zhou, Y.S. Park, Y.H. Kim, Y. Kim and K.P. Kim, “Binary matrix for MALDI imaging mass spectrometry of phospholipids in both ion modes” Anal. Chem. 83, 1252 (2011). doi: http://dx.doi.org/10.1021/ac1029659
- Z. Guo and L. He, “A binary matrix for background suppression in MALDI-MS of small molecules”, Anal. Bioanal. Chem. 387, 1939 (2007). doi: http://dx.doi.org/10.1007/s00216-006-1100-3
- J. Hong, T. Kim and J. Kim, “Tertiary matrices for the analysis of polyethylene glycols using MALDI-TOF MS”, Mass Spectrom. Lett. 5, 49 (2014). doi: http://dx.doi.org/10.5478/MSL.2014.5.2.49
- J. Kemptner, M. Marchetti-Deschmann, R. Mach, I.S. Druzhinina, C.P. Kubicek and G. Allmaier, “Evaluation of matrix-assisted laser desorption/ionization (MALDI) preparation techniques for surface characterization of intact Fusarium spores by MALDI linear time-of-flight mass spectrometry”, Rapid Commun. Mass Spectrom. 23, 877 (2009). doi: http://dx.doi.org/10.1002/rcm.3949
- J. Chalupová, M. Sedlárˇová, M. Helmel, P. Rˇehulka, M. Marchetti-Deschmann, G. Allmaier and M. Šebela, “MALDI-based intact spore mass spectrometry of downy and powdery mildews”, J. Mass Spectrom. 47, 978 (2012). doi: http://dx.doi.org/10.1002/jms.3046